2´-O-Methyl (2´-OMe) Panels
2´-O-Methyl (2´-OMe) Panels
ModDetect™ 2′-O-Methyl (2′-OMe) Panels are designed for the detection of 2′-OMe chemical modifications independent of sequence or position. These reagents can be applied across multiple assay formats to support studies of distribution, uptake, and localization of chemically modified oligonucleotides.
The Role of 2´-OMe Modifications in ASO Drug Design
2′-O-Methyl (2′-OMe) chemical modification of oligonucleotides is a second-generation modification often incorporated into antisense oligonucleotides (ASOs), small interfering RNA (siRNA), aptamers, and messenger RNA (mRNA). OMe modification enhances stability, minimizes off-target binding, and reduces innate immune response, which are key benefits when developing safer, more effective RNA therapeutics. However, excessive incorporation of OMe modification may lower the potency of oligonucleotide drugs, requiring consideration for both pros and cons in drug design.
ModDetect™ 2’-OMe panels feature highly specific antibodies that recognize the OMe chemical modification, detecting nucleic acid drugs independent of sequence. These antibodies support quantitative immunoassays across various modalities (ASOs, siRNA, aptamers, mRNA) and can be used to localize and quantify siRNA strands synthesized with distinct chemical modifications (i.e. when phosphorothioate (PS) is present on both strands, but with MOE on the sense strand and OMe on the antisense strand.)
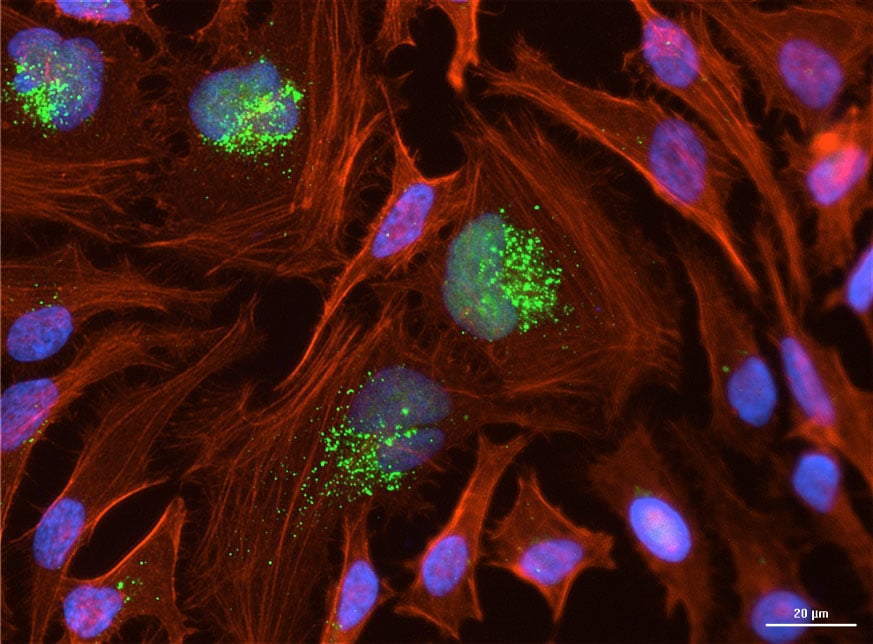
Intracellular Localization of 2′-OMe–Modified siRNA
ModDetect™ Anti-2′-OMe antibodies enable clear visualization of 2′-O-methyl–modified siRNA within cells, providing spatial insight into intracellular distribution.
In HeLa cells, lumasiran siRNA was detected using clone OME3, with cytoplasmic staining observed in a pattern consistent with endosomal sequestration. Co-staining with phalloidin and DAPI provided structural context, confirming cellular localization relative to cytoskeletal features and nuclei. These data support the use of ModDetect™ for assessing cellular uptake and intracellular positioning of OMe-modified oligonucleotides.
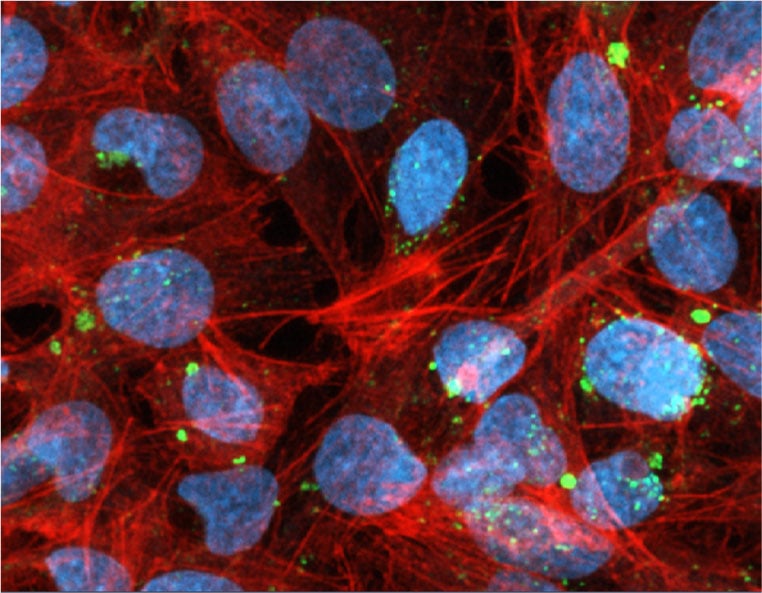
Cytoplasmic Distribution of 2′-OMe–Modified siRNA
ModDetect™ Anti-2′-OMe antibodies enable visualization of 2′-O-methyl–modified siRNA in physiologically relevant cell models.
In Hep G2 cells, lumasiran siRNA was detected using clone OME3, with punctate staining observed within the cytoplasmic consistent with entrapment. Co-staining with phalloidin and DAPI provided structural context, confirming localization relative to the actin cytoskeleton and nuclei. These results demonstrate the utility of ModDetect™ for evaluating intracellular distribution of OMe-modified oligonucleotides in human derived liver cancer cells (hepatoblastoma).
Available ModDetect™ Panels and Clones
The 2′-OMe ModDetect™ Panels include a set of unconjugated or biotinylated monoclonal antibodies validated for the detection of 2′-O-methyl–modified antisense oligonucleotides. Individual antibody clones are also available for researchers who require flexibility in assay development or wish to evaluate clone-specific performance.
| 2′-OMe Panels | Format |
| ModDetect™ 2′-O-Methyl (2′-OMe) Panel | Unconjugated |
| ModDetect™ 2′-O-Methyl (2′-OMe) Biotinylated Panel | Biotin |
| 2′-OMe Clones | Included in Panel |
| Unconjugated Clone OME1 | ✓ |
| Unconjugated Clone OME2 | ✓ |
| Unconjugated Clone OME3 | ✓ |
| Unconjugated Clone OME4 | ✓ |
| Unconjugated Clone OME5 | ✓ |
| Biotin Clone OME1 | ✓ |
| Biotin Clone OME2 | ✓ |
| Biotin Clone OME3 | ✓ |
| Biotin Clone OME4 | ✓ |
| Biotin Clone OME5 | ✓ |